The Loom format is an HDF5-based
file format used by loompy and RNA velocity tools (velocyto, scVelo).
scConvert provides readLoom() and writeLoom()
for round-trip conversion between Seurat objects and Loom files with no
external dependencies.
Read a Loom file
scConvert ships a 500-cell PBMC dataset in Loom format.
readLoom() loads it directly into a Seurat object.
loom_file <- system.file("extdata", "pbmc_demo.loom", package = "scConvert")
pbmc <- readLoom(loom_file)
pbmc
#> An object of class Seurat
#> 2000 features across 500 samples within 1 assay
#> Active assay: RNA (2000 features, 0 variable features)
#> 2 layers present: counts, data
#> 1 dimensional reduction calculated: umapThe reader reconstructs cell metadata, gene metadata, and dimensional reductions from the Loom column and row attributes:
cat("Cells:", ncol(pbmc), "\n")
#> Cells: 500
cat("Genes:", nrow(pbmc), "\n")
#> Genes: 2000
cat("Reductions:", paste(names(pbmc@reductions), collapse = ", "), "\n")
#> Reductions: umap
cat("Metadata columns:", paste(colnames(pbmc[[]]), collapse = ", "), "\n")
#> Metadata columns: orig.ident, nCount_RNA, nFeature_RNA, RNA_snn_res.0.5, percent.mt, seurat_annotations, seurat_clustersThe nine annotated cell types are preserved:
DimPlot(pbmc, reduction = "umap", group.by = "seurat_annotations",
label = TRUE, pt.size = 0.5) + NoLegend()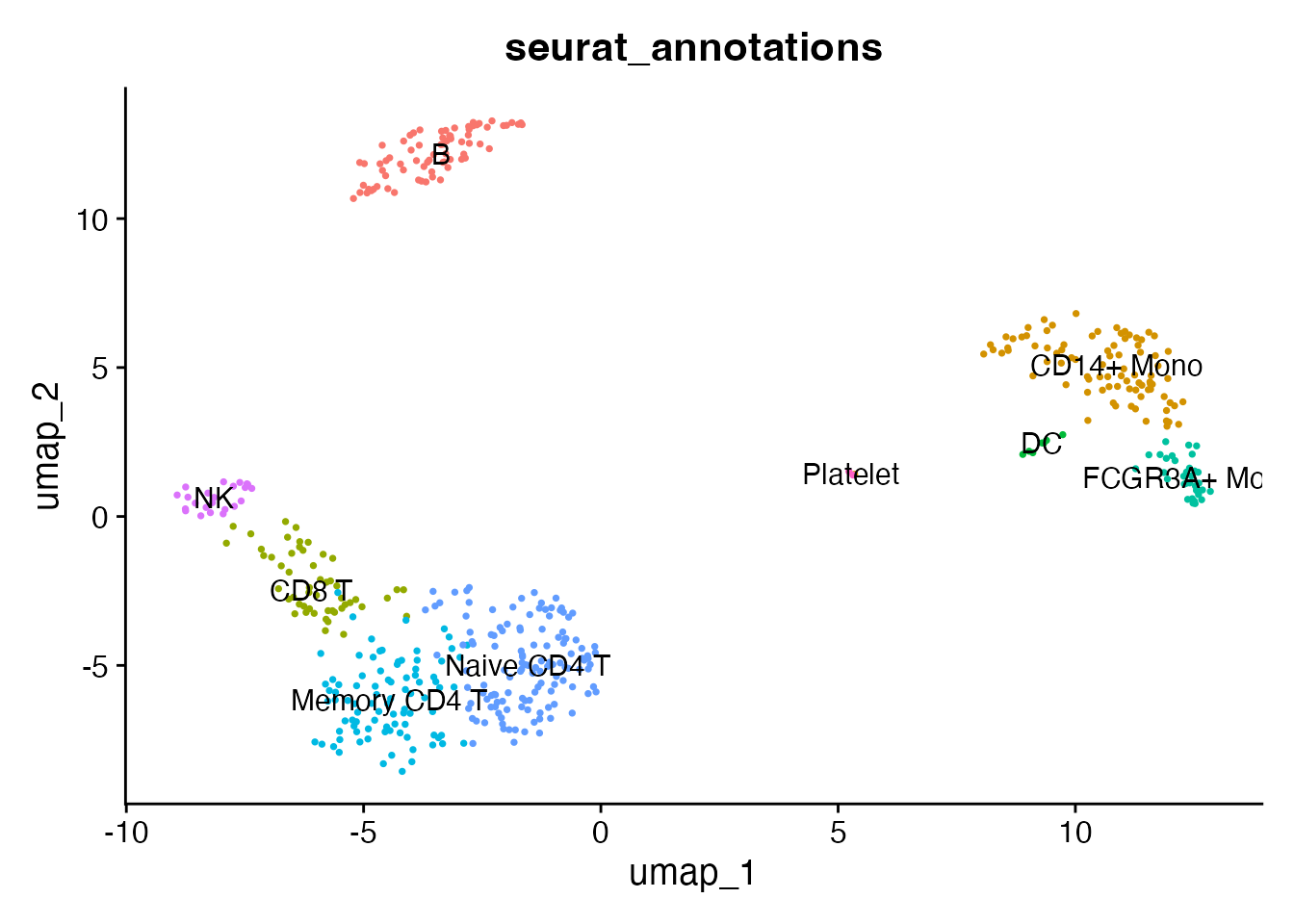
Expression data is fully available. LYZ is a monocyte marker:
FeaturePlot(pbmc, features = "LYZ", pt.size = 0.5)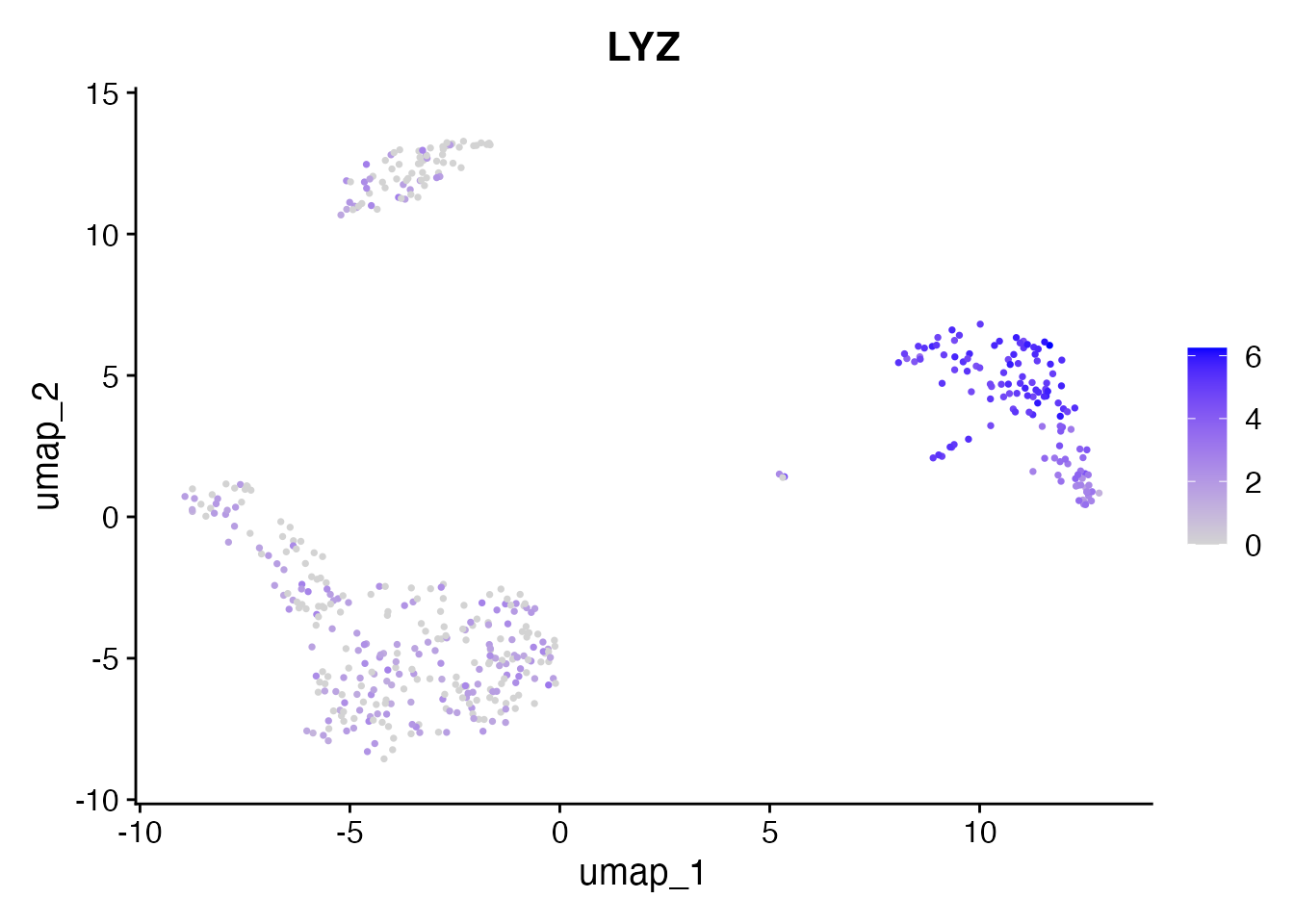
Write a Seurat object to Loom
writeLoom() saves the default assay’s expression data as
the main matrix, with cell metadata as column attributes and gene
metadata as row attributes. Dimensional reductions are stored as
additional column attributes.
Verify the round-trip
Read the written Loom file back and compare the UMAP projections.
pbmc_rt <- readLoom(loom_path)
library(patchwork)
p1 <- DimPlot(pbmc_seurat, reduction = "umap", group.by = "seurat_annotations",
label = TRUE, pt.size = 0.5) + NoLegend() + ggtitle("Original (.rds)")
p2 <- DimPlot(pbmc_rt, reduction = "umap", group.by = "seurat_annotations",
label = TRUE, pt.size = 0.5) + NoLegend() + ggtitle("Round-trip (.loom)")
p1 + p2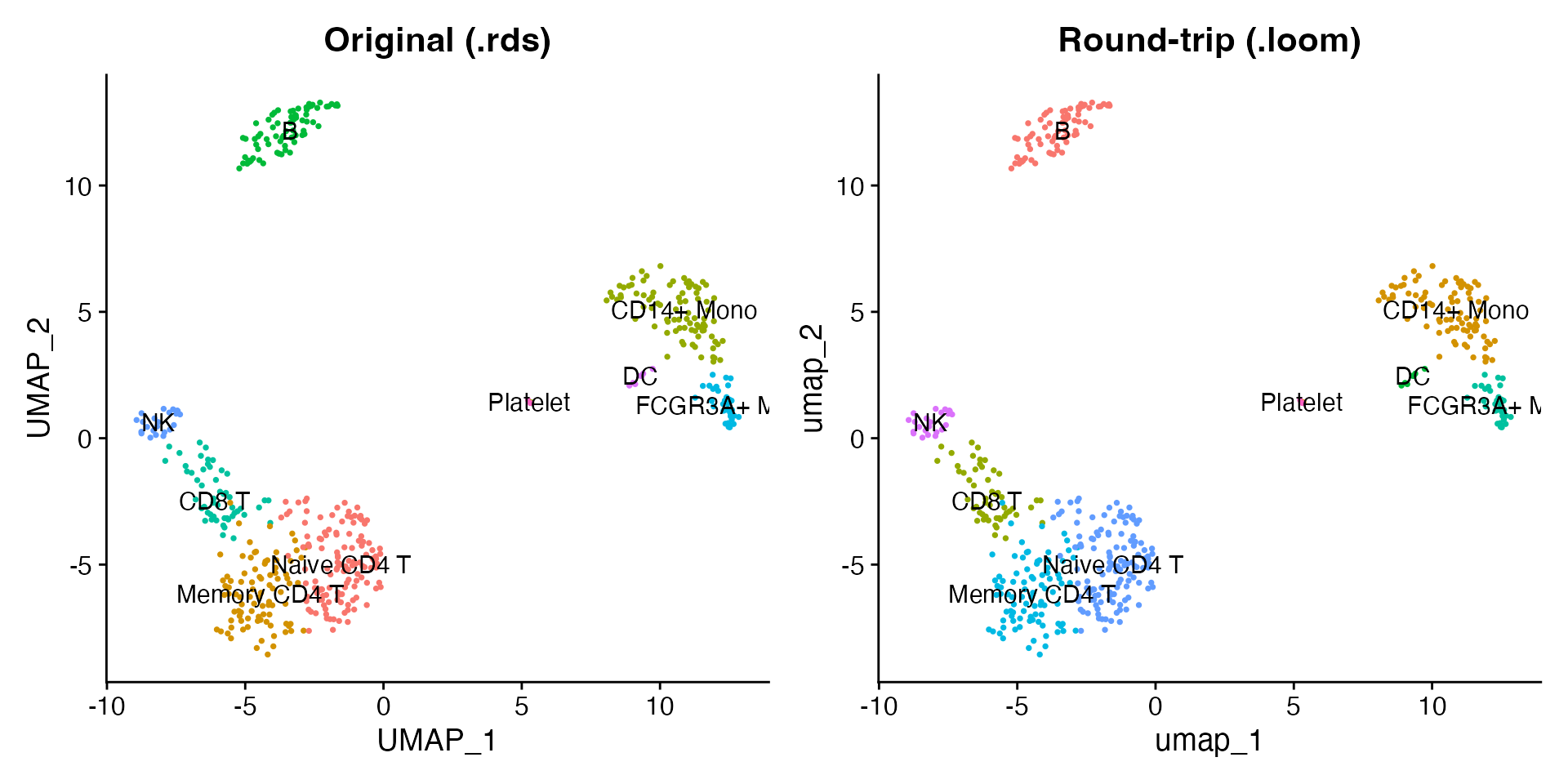
What is preserved
Loom is a simpler format than h5ad or h5Seurat. Here is a summary of what round-trips and what does not:
| Component | Preserved? | Notes |
|---|---|---|
| Expression matrix | Yes | Stored as /matrix
|
| Raw counts | Yes | Stored in /layers/counts
|
| Cell metadata | Yes | Each column becomes a /col_attrs entry |
| Gene metadata | Yes | Each column becomes a /row_attrs entry |
| PCA / UMAP embeddings | Yes | Stored as column attributes |
| Nearest-neighbor graphs | No | Not native to Loom; recompute with FindNeighbors()
|
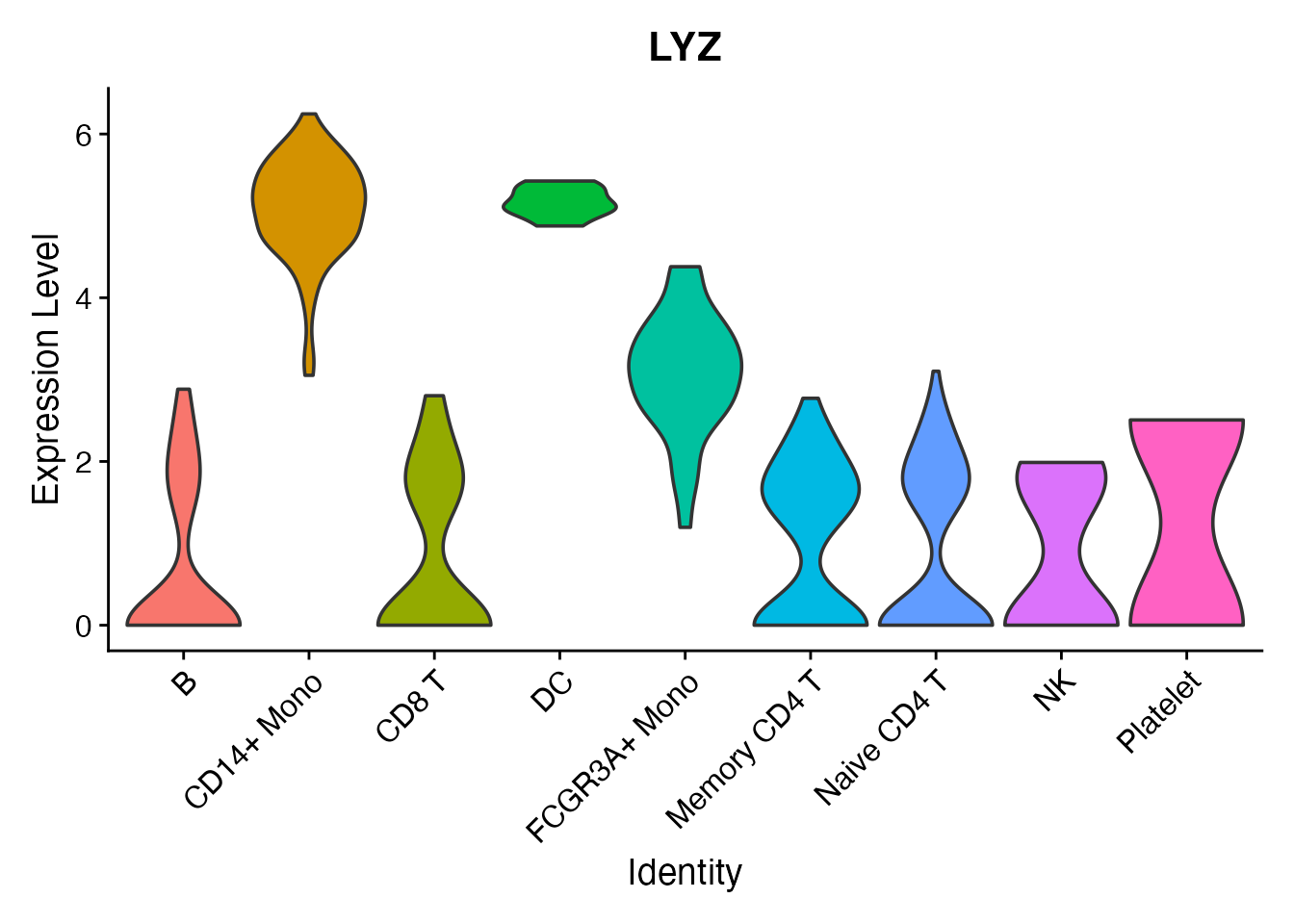
Per-cluster expression
A violin plot confirms that per-cluster expression distributions are preserved through Loom conversion:
Python interop (optional)
The Loom files produced by writeLoom() are compatible
with loompy, scanpy, and scVelo.
Clean up
unlink(loom_path)