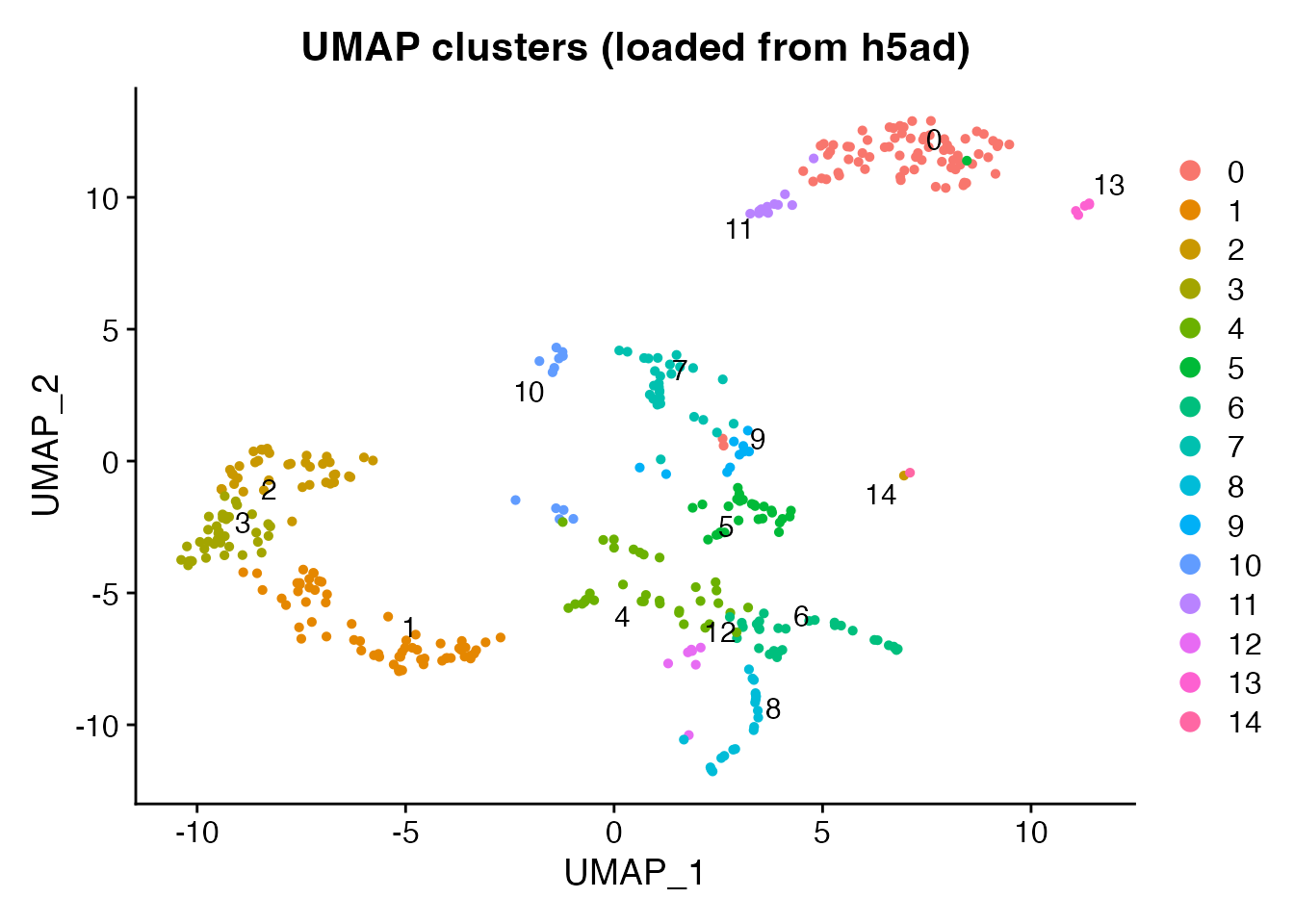
Introduction
10x Genomics Visium captures spatially-resolved gene expression across tissue sections. When converting Visium data between Seurat and h5ad, scConvert preserves spot coordinates, tissue images, scale factors, and all associated metadata so the data is immediately usable in both R (Seurat) and Python (scanpy/squidpy) ecosystems.
Read spatial data from h5ad
The shipped spatial_demo.h5ad contains 400 mouse brain
Visium spots with 1,500 genes, PCA/UMAP reductions, and cluster labels.
readH5AD() rebuilds the full Seurat spatial object,
including the tissue image.
h5ad_file <- system.file("extdata", "spatial_demo.h5ad", package = "scConvert")
brain <- readH5AD(h5ad_file, verbose = FALSE)
cat("Spots:", ncol(brain), "\n")
#> Spots: 400
cat("Genes:", nrow(brain), "\n")
#> Genes: 1500
cat("Image:", paste(names(brain@images), collapse = ", "), "\n")
#> Image: anterior1
cat("Assay:", paste(names(brain@assays), collapse = ", "), "\n")
#> Assay: RNASpatial expression of a brain marker
Hpca (Hippocalcin) marks hippocampal neurons and shows clear regional expression in the mouse brain.
SpatialFeaturePlot(brain, features = "Hpca", pt.size.factor = 1.6) +
ggtitle("Hpca expression (loaded from h5ad)")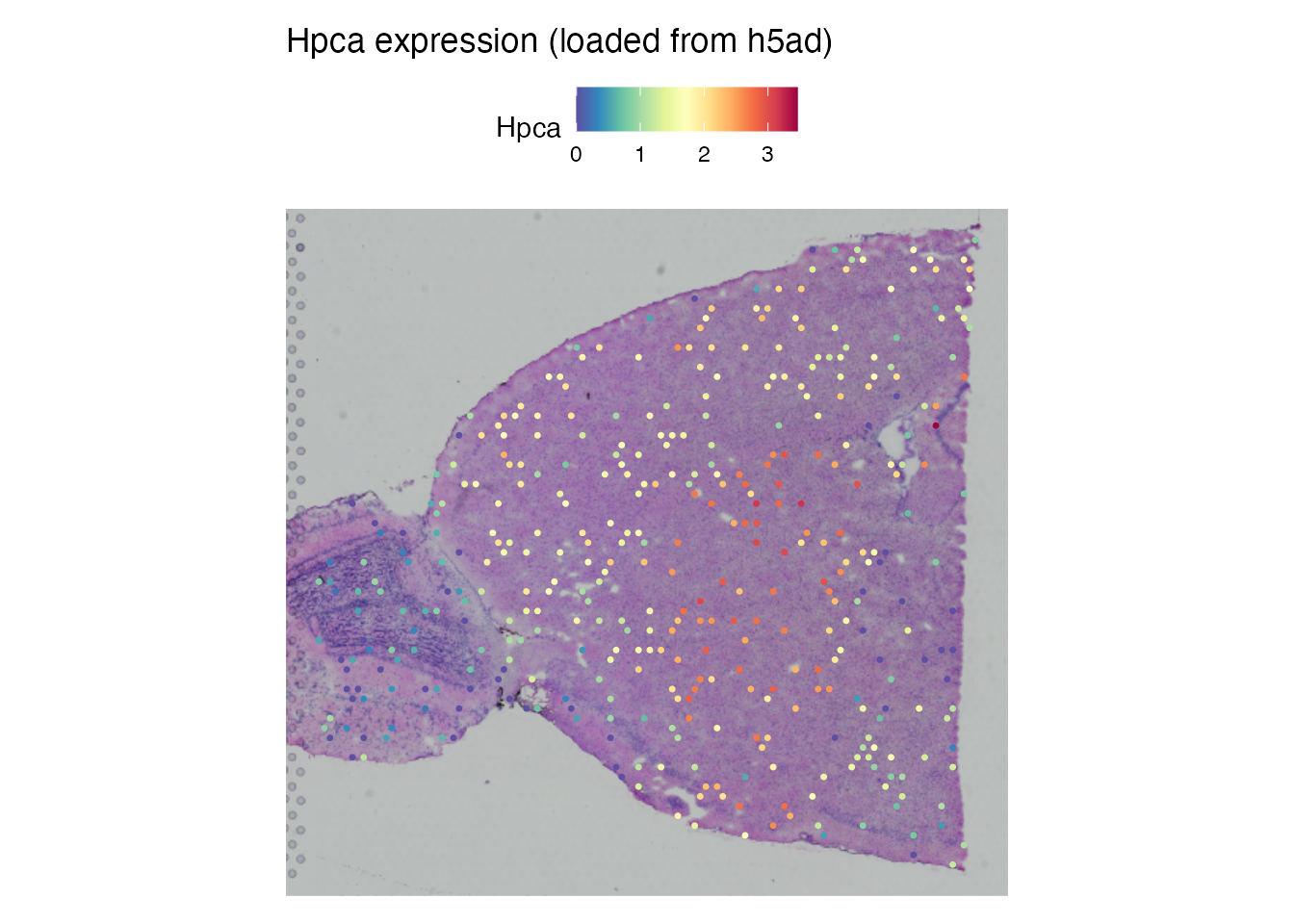
Write-and-read roundtrip
Write the spatial object to a new h5ad file, read it back, and confirm that coordinates, images, scale factors, and expression values are all preserved.
h5ad_rt <- file.path(tempdir(), "spatial_roundtrip.h5ad")
writeH5AD(brain, h5ad_rt, overwrite = TRUE)
cat("Wrote:", round(file.size(h5ad_rt) / 1024^2, 1), "MB\n")
#> Wrote: 3.2 MB
brain_rt <- readH5AD(h5ad_rt, verbose = FALSE)
cat("Roundtrip spots:", ncol(brain_rt), "\n")
#> Roundtrip spots: 400
cat("Roundtrip genes:", nrow(brain_rt), "\n")
#> Roundtrip genes: 1500
cat("Image:", paste(names(brain_rt@images), collapse = ", "), "\n")
#> Image: anterior1Compare spatial plots
The spatial expression pattern should be identical after roundtrip.
SpatialFeaturePlot(brain_rt, features = "Hpca", pt.size.factor = 1.6) +
ggtitle("Hpca expression (after h5ad roundtrip)")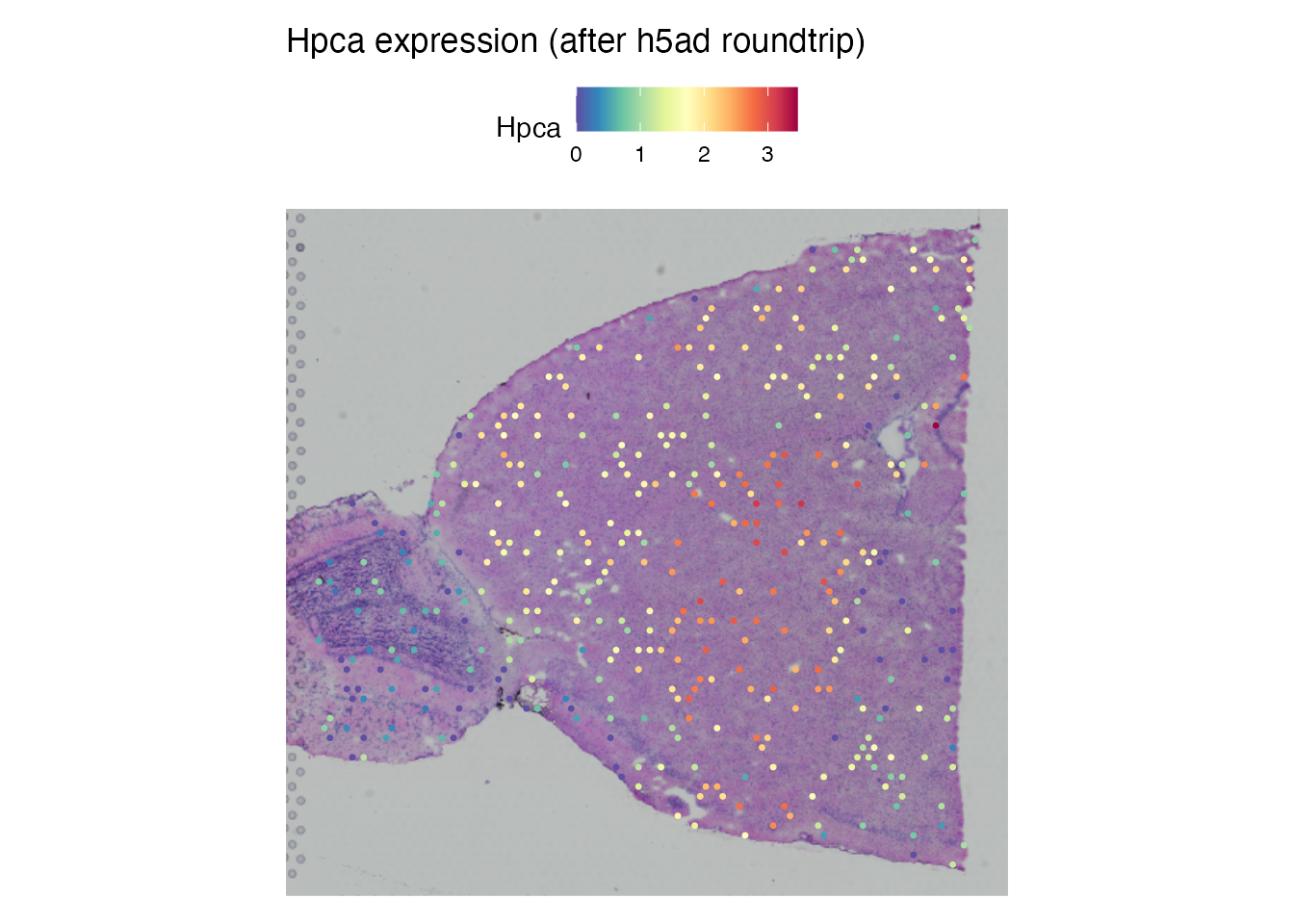
Verify data integrity
cat("Dimensions match:",
ncol(brain) == ncol(brain_rt) && nrow(brain) == nrow(brain_rt), "\n")
#> Dimensions match: TRUE
cat("Barcodes match:", all(colnames(brain) == colnames(brain_rt)), "\n")
#> Barcodes match: TRUE
cat("Clusters match:",
all(as.character(brain$seurat_clusters) ==
as.character(brain_rt$seurat_clusters)), "\n")
#> Clusters match: TRUEWhat gets preserved
scConvert maintains full fidelity for Visium spatial data during conversion:
| Component | h5ad location | Preserved |
|---|---|---|
| Expression matrix |
X / raw/X
|
Yes |
| Spot coordinates | obsm/spatial |
Yes |
| Tissue image | uns/spatial/*/images |
Yes |
| Scale factors | uns/spatial/*/scalefactors |
Yes |
| Cell metadata | obs |
Yes |
| Reductions (PCA, UMAP) | obsm |
Yes |
Use in Python (scanpy/squidpy)
The exported h5ad works directly with scanpy and squidpy. Images, coordinates, and scale factors use standard scanpy conventions.
# Requires Python with scanpy and squidpy installed
import scanpy as sc
adata = sc.read_h5ad("spatial_demo.h5ad")
print(adata)
print(f"Spatial coords shape: {adata.obsm['spatial'].shape}")
# Spatial scatter plot
sc.pl.spatial(adata, color="seurat_clusters")
# Squidpy spatial analysis
import squidpy as sq
sq.gr.spatial_neighbors(adata, coord_type="generic")
sq.gr.spatial_autocorr(adata, mode="moran", genes=["Hpca", "Ttr"])Session Info
sessionInfo()
#> R version 4.6.0 (2026-04-24)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.3 Seurat_5.5.0 SeuratObject_5.4.0 sp_2.2-1
#> [5] scConvert_0.2.0
#>
#> loaded via a namespace (and not attached):
#> [1] RColorBrewer_1.1-3 jsonlite_2.0.0 magrittr_2.0.5
#> [4] spatstat.utils_3.2-2 farver_2.1.2 rmarkdown_2.31
#> [7] fs_2.1.0 ragg_1.5.2 vctrs_0.7.3
#> [10] ROCR_1.0-12 spatstat.explore_3.8-0 htmltools_0.5.9
#> [13] sass_0.4.10 sctransform_0.4.3 parallelly_1.47.0
#> [16] KernSmooth_2.23-26 bslib_0.10.0 htmlwidgets_1.6.4
#> [19] desc_1.4.3 ica_1.0-3 plyr_1.8.9
#> [22] plotly_4.12.0 zoo_1.8-15 cachem_1.1.0
#> [25] igraph_2.3.0 mime_0.13 lifecycle_1.0.5
#> [28] pkgconfig_2.0.3 Matrix_1.7-5 R6_2.6.1
#> [31] fastmap_1.2.0 MatrixGenerics_1.24.0 fitdistrplus_1.2-6
#> [34] future_1.70.0 shiny_1.13.0 digest_0.6.39
#> [37] S4Vectors_0.50.0 patchwork_1.3.2 tensor_1.5.1
#> [40] RSpectra_0.16-2 irlba_2.3.7 GenomicRanges_1.64.0
#> [43] textshaping_1.0.5 labeling_0.4.3 progressr_0.19.0
#> [46] spatstat.sparse_3.1-0 httr_1.4.8 polyclip_1.10-7
#> [49] abind_1.4-8 compiler_4.6.0 bit64_4.8.0
#> [52] withr_3.0.2 S7_0.2.2 fastDummies_1.7.6
#> [55] MASS_7.3-65 tools_4.6.0 lmtest_0.9-40
#> [58] otel_0.2.0 httpuv_1.6.17 future.apply_1.20.2
#> [61] goftest_1.2-3 glue_1.8.1 nlme_3.1-169
#> [64] promises_1.5.0 grid_4.6.0 Rtsne_0.17
#> [67] cluster_2.1.8.2 reshape2_1.4.5 generics_0.1.4
#> [70] hdf5r_1.3.12 gtable_0.3.6 spatstat.data_3.1-9
#> [73] tidyr_1.3.2 data.table_1.18.2.1 BiocGenerics_0.58.0
#> [76] BPCells_0.3.1 spatstat.geom_3.7-3 RcppAnnoy_0.0.23
#> [79] ggrepel_0.9.8 RANN_2.6.2 pillar_1.11.1
#> [82] stringr_1.6.0 spam_2.11-3 RcppHNSW_0.6.0
#> [85] later_1.4.8 splines_4.6.0 dplyr_1.2.1
#> [88] lattice_0.22-9 survival_3.8-6 bit_4.6.0
#> [91] deldir_2.0-4 tidyselect_1.2.1 miniUI_0.1.2
#> [94] pbapply_1.7-4 knitr_1.51 gridExtra_2.3
#> [97] Seqinfo_1.2.0 IRanges_2.46.0 scattermore_1.2
#> [100] stats4_4.6.0 xfun_0.57 matrixStats_1.5.0
#> [103] stringi_1.8.7 lazyeval_0.2.3 yaml_2.3.12
#> [106] evaluate_1.0.5 codetools_0.2-20 tibble_3.3.1
#> [109] cli_3.6.6 uwot_0.2.4 xtable_1.8-8
#> [112] reticulate_1.46.0 systemfonts_1.3.2 jquerylib_0.1.4
#> [115] Rcpp_1.1.1-1.1 globals_0.19.1 spatstat.random_3.4-5
#> [118] png_0.1-9 spatstat.univar_3.1-7 parallel_4.6.0
#> [121] pkgdown_2.2.0 dotCall64_1.2 listenv_0.10.1
#> [124] viridisLite_0.4.3 scales_1.4.0 ggridges_0.5.7
#> [127] purrr_1.2.2 crayon_1.5.3 rlang_1.2.0
#> [130] cowplot_1.2.0