Introduction
SpatialData is the
scverse standard for spatial omics data. A
.spatialdata.zarr store combines expression tables, spatial
coordinates, shapes, and tissue images in a single Zarr directory.
Technologies like Visium, MERFISH, Xenium, Slide-seq, and CODEX all have
SpatialData representations, making it a unifying format for the spatial
transcriptomics ecosystem.
scConvert provides readSpatialData() and
writeSpatialData() to move between SpatialData stores and
Seurat objects, with no Python dependency.
Read spatial data from h5ad
We start by reading a 400-spot Visium mouse brain dataset shipped with scConvert as an h5ad file. This dataset includes spatial coordinates, a tissue image, PCA/UMAP embeddings, and 15 clusters across 1500 genes.
h5ad_path <- system.file("extdata", "spatial_demo.h5ad", package = "scConvert")
brain <- readH5AD(h5ad_path, verbose = FALSE)
brain
#> An object of class Seurat
#> 1500 features across 400 samples within 1 assay
#> Active assay: RNA (1500 features, 1500 variable features)
#> 2 layers present: counts, data
#> 2 dimensional reductions calculated: pca, umap
#> 1 spatial field of view present: anterior1
SpatialFeaturePlot(brain, features = "Ttr") +
ggtitle("Ttr expression -- Visium mouse brain (from h5ad)")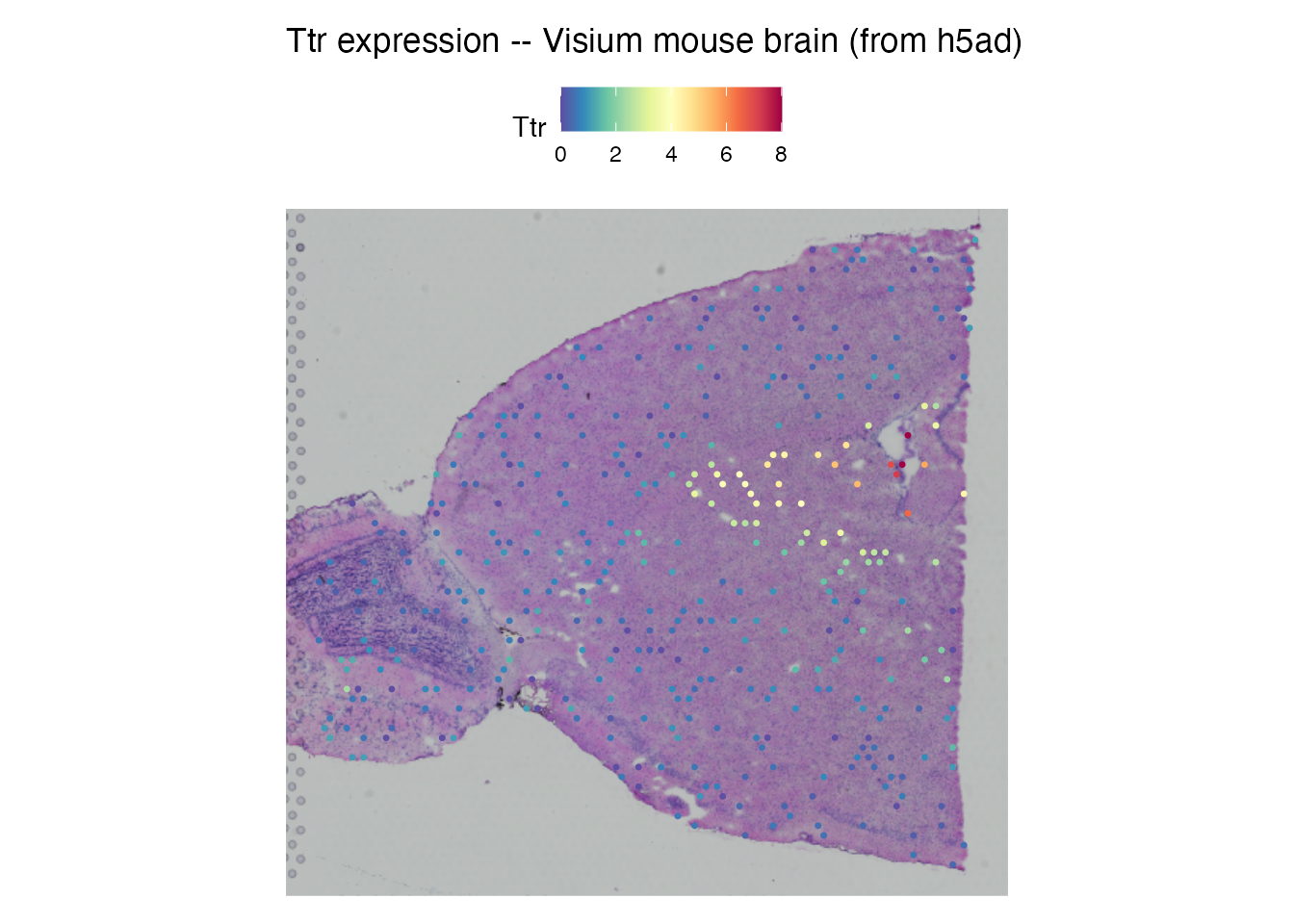
Ttr (transthyretin) marks the choroid plexus. The spatial expression pattern confirms the data was loaded correctly from the h5ad file.
DimPlot(brain, reduction = "umap", label = TRUE, pt.size = 1) +
ggtitle("Cluster assignments (from h5ad)") + NoLegend()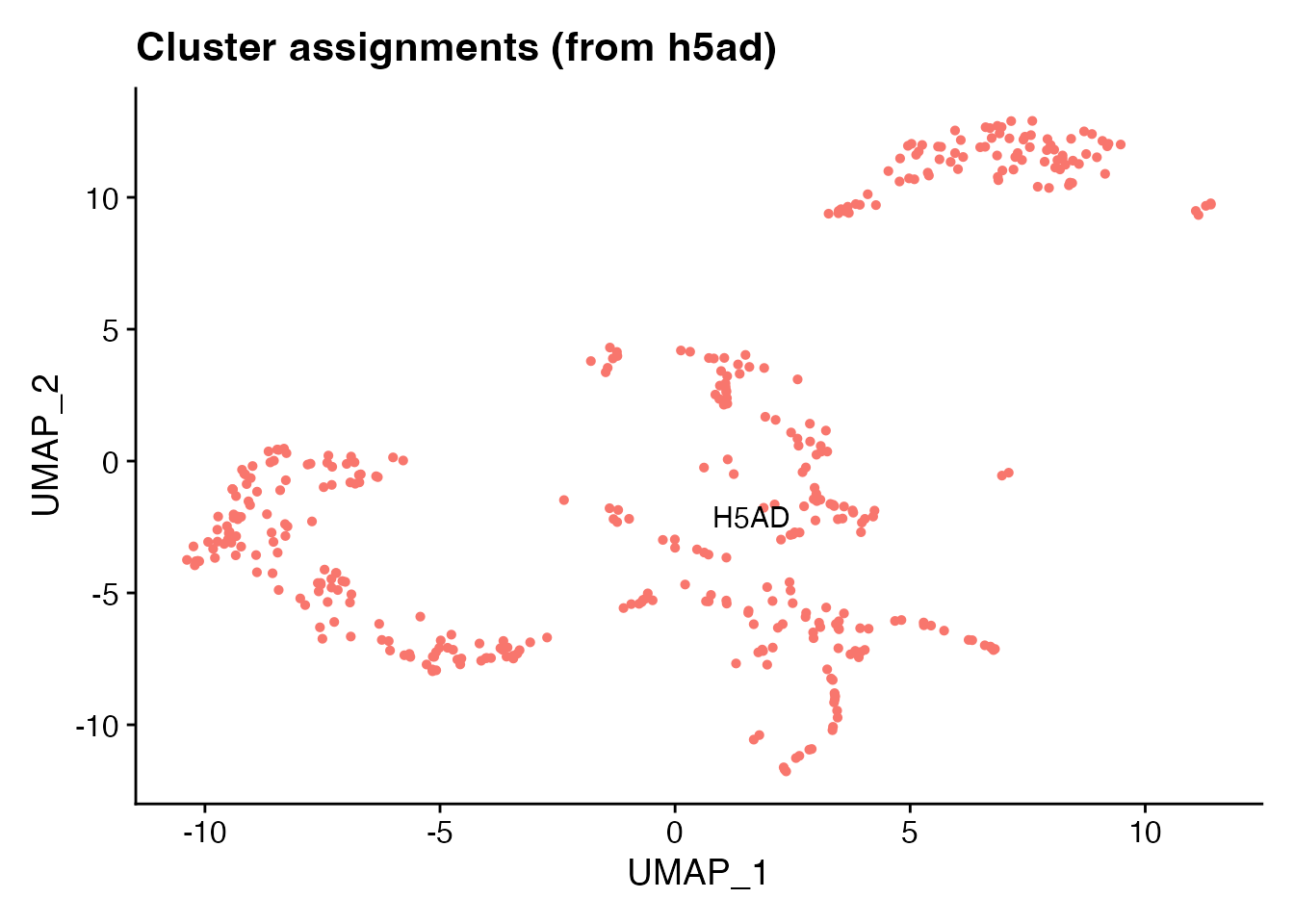
Write to SpatialData and read back
SpatialData stores have a complex directory structure with separate groups for tables, shapes, images, and coordinate systems. We write the Seurat object to a SpatialData Zarr store and read it back to verify the round-trip.
sd_path <- file.path(tempdir(), "brain.spatialdata.zarr")
writeSpatialData(brain, sd_path, overwrite = TRUE, verbose = FALSE)
cat("SpatialData store written to:", sd_path, "\n")
#> SpatialData store written to: /tmp/RtmpsLVqa3/brain.spatialdata.zarr
brain_rt <- readSpatialData(sd_path, verbose = FALSE)
cat("Cells:", ncol(brain_rt), "| Genes:", nrow(brain_rt), "\n")
#> Cells: 400 | Genes: 1500
cat("Reductions:", paste(names(brain_rt@reductions), collapse = ", "), "\n")
#> Reductions: pca, umap
cat("Images:", paste(names(brain_rt@images), collapse = ", "), "\n")
#> Images: anterior1Compare original and round-trip
library(patchwork)
p1 <- FeaturePlot(brain, features = "Ttr", pt.size = 1) +
ggtitle("Ttr -- Original (h5ad)")
p2 <- FeaturePlot(brain_rt, features = "Ttr", pt.size = 1) +
ggtitle("Ttr -- After SpatialData round-trip")
p1 + p2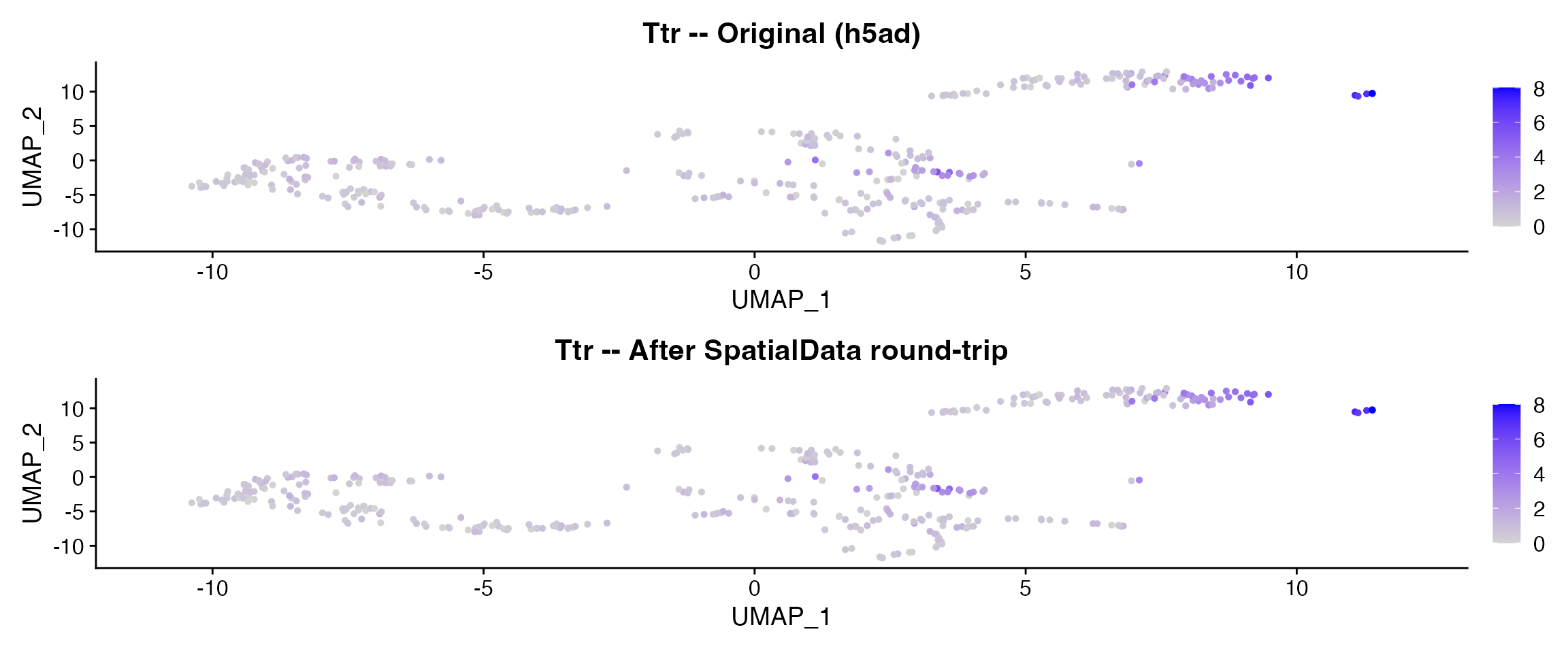
Direct pair converters
Convert between SpatialData and other formats in a single call:
# SpatialData <-> h5ad
SpatialDataToH5AD("sample.spatialdata.zarr", "sample.h5ad")
H5ADToSpatialData("sample.h5ad", "sample.spatialdata.zarr")
# SpatialData <-> h5Seurat
SpatialDataToH5Seurat("sample.spatialdata.zarr", "sample.h5seurat")
H5SeuratToSpatialData("sample.h5seurat", "sample.spatialdata.zarr")
# Or use the universal dispatcher
scConvert("sample.spatialdata.zarr", dest = "sample.h5ad", overwrite = TRUE)Limitations
-
Images: Large OME-NGFF images (>100M pixels) are
skipped to avoid memory issues. Use
images = FALSEto read coordinates only. - Coordinate transforms: SpatialData supports affine transformations between elements. These are not currently applied during reading; coordinates are used as-is.
- Labels/segmentation: Cell and nucleus segmentation masks are recorded by name but not loaded (they have no natural Seurat representation).
-
Multiple tables: Only one table is read per call.
Use the
tableargument to select which one.
Python interoperability
SpatialData stores written by scConvert are readable by the Python spatialdata library. Requires Python with spatialdata installed.
Session Info
sessionInfo()
#> R version 4.6.0 (2026-04-24)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] patchwork_1.3.2 ggplot2_4.0.3 Seurat_5.5.0 SeuratObject_5.4.0
#> [5] sp_2.2-1 scConvert_0.2.0
#>
#> loaded via a namespace (and not attached):
#> [1] deldir_2.0-4 pbapply_1.7-4 gridExtra_2.3
#> [4] rlang_1.2.0 magrittr_2.0.5 RcppAnnoy_0.0.23
#> [7] otel_0.2.0 spatstat.geom_3.7-3 matrixStats_1.5.0
#> [10] ggridges_0.5.7 compiler_4.6.0 png_0.1-9
#> [13] systemfonts_1.3.2 vctrs_0.7.3 reshape2_1.4.5
#> [16] hdf5r_1.3.12 stringr_1.6.0 crayon_1.5.3
#> [19] pkgconfig_2.0.3 fastmap_1.2.0 labeling_0.4.3
#> [22] promises_1.5.0 rmarkdown_2.31 ragg_1.5.2
#> [25] bit_4.6.0 purrr_1.2.2 xfun_0.57
#> [28] cachem_1.1.0 jsonlite_2.0.0 goftest_1.2-3
#> [31] later_1.4.8 spatstat.utils_3.2-2 irlba_2.3.7
#> [34] parallel_4.6.0 cluster_2.1.8.2 R6_2.6.1
#> [37] ica_1.0-3 spatstat.data_3.1-9 bslib_0.10.0
#> [40] stringi_1.8.7 RColorBrewer_1.1-3 reticulate_1.46.0
#> [43] spatstat.univar_3.1-7 parallelly_1.47.0 lmtest_0.9-40
#> [46] jquerylib_0.1.4 scattermore_1.2 Rcpp_1.1.1-1.1
#> [49] knitr_1.51 tensor_1.5.1 future.apply_1.20.2
#> [52] zoo_1.8-15 sctransform_0.4.3 httpuv_1.6.17
#> [55] Matrix_1.7-5 splines_4.6.0 igraph_2.3.0
#> [58] tidyselect_1.2.1 abind_1.4-8 yaml_2.3.12
#> [61] spatstat.random_3.4-5 spatstat.explore_3.8-0 codetools_0.2-20
#> [64] miniUI_0.1.2 listenv_0.10.1 plyr_1.8.9
#> [67] lattice_0.22-9 tibble_3.3.1 withr_3.0.2
#> [70] shiny_1.13.0 S7_0.2.2 ROCR_1.0-12
#> [73] evaluate_1.0.5 Rtsne_0.17 future_1.70.0
#> [76] fastDummies_1.7.6 desc_1.4.3 survival_3.8-6
#> [79] polyclip_1.10-7 fitdistrplus_1.2-6 pillar_1.11.1
#> [82] KernSmooth_2.23-26 plotly_4.12.0 generics_0.1.4
#> [85] RcppHNSW_0.6.0 scales_1.4.0 globals_0.19.1
#> [88] xtable_1.8-8 glue_1.8.1 lazyeval_0.2.3
#> [91] tools_4.6.0 data.table_1.18.2.1 RSpectra_0.16-2
#> [94] RANN_2.6.2 fs_2.1.0 dotCall64_1.2
#> [97] cowplot_1.2.0 grid_4.6.0 tidyr_1.3.2
#> [100] nlme_3.1-169 cli_3.6.6 spatstat.sparse_3.1-0
#> [103] textshaping_1.0.5 spam_2.11-3 viridisLite_0.4.3
#> [106] dplyr_1.2.1 uwot_0.2.4 gtable_0.3.6
#> [109] sass_0.4.10 digest_0.6.39 progressr_0.19.0
#> [112] ggrepel_0.9.8 htmlwidgets_1.6.4 farver_2.1.2
#> [115] htmltools_0.5.9 pkgdown_2.2.0 lifecycle_1.0.5
#> [118] httr_1.4.8 mime_0.13 bit64_4.8.0
#> [121] MASS_7.3-65