Introduction
Single-cell ATAC-seq measures chromatin accessibility at single-cell resolution. The core data structure is a peak-by-cell count matrix – structurally identical to a gene-by-cell RNA matrix. scConvert converts this matrix along with cell metadata, embeddings, and cluster labels between formats, enabling interoperability between R (Seurat/Signac) and Python (scanpy/episcanpy) workflows.
This vignette demonstrates the h5ad roundtrip using the shipped PBMC demo data as a stand-in. The matrix structure and conversion mechanics are the same regardless of whether rows represent genes or peaks.
Load demo data
obj <- readRDS(system.file("extdata", "pbmc_demo.rds", package = "scConvert"))
cat("Cells:", ncol(obj), "\n")
#> Cells: 500
cat("Features:", nrow(obj), "\n")
#> Features: 2000
cat("Reductions:", paste(names(obj@reductions), collapse = ", "), "\n")
#> Reductions: pca, umap
cat("Metadata columns:", paste(colnames(obj[[]]), collapse = ", "), "\n")
#> Metadata columns: orig.ident, nCount_RNA, nFeature_RNA, seurat_annotations, percent.mt, RNA_snn_res.0.5, seurat_clusters
DimPlot(obj, group.by = "seurat_annotations", label = TRUE, pt.size = 0.8) +
ggtitle("PBMC clusters (original)")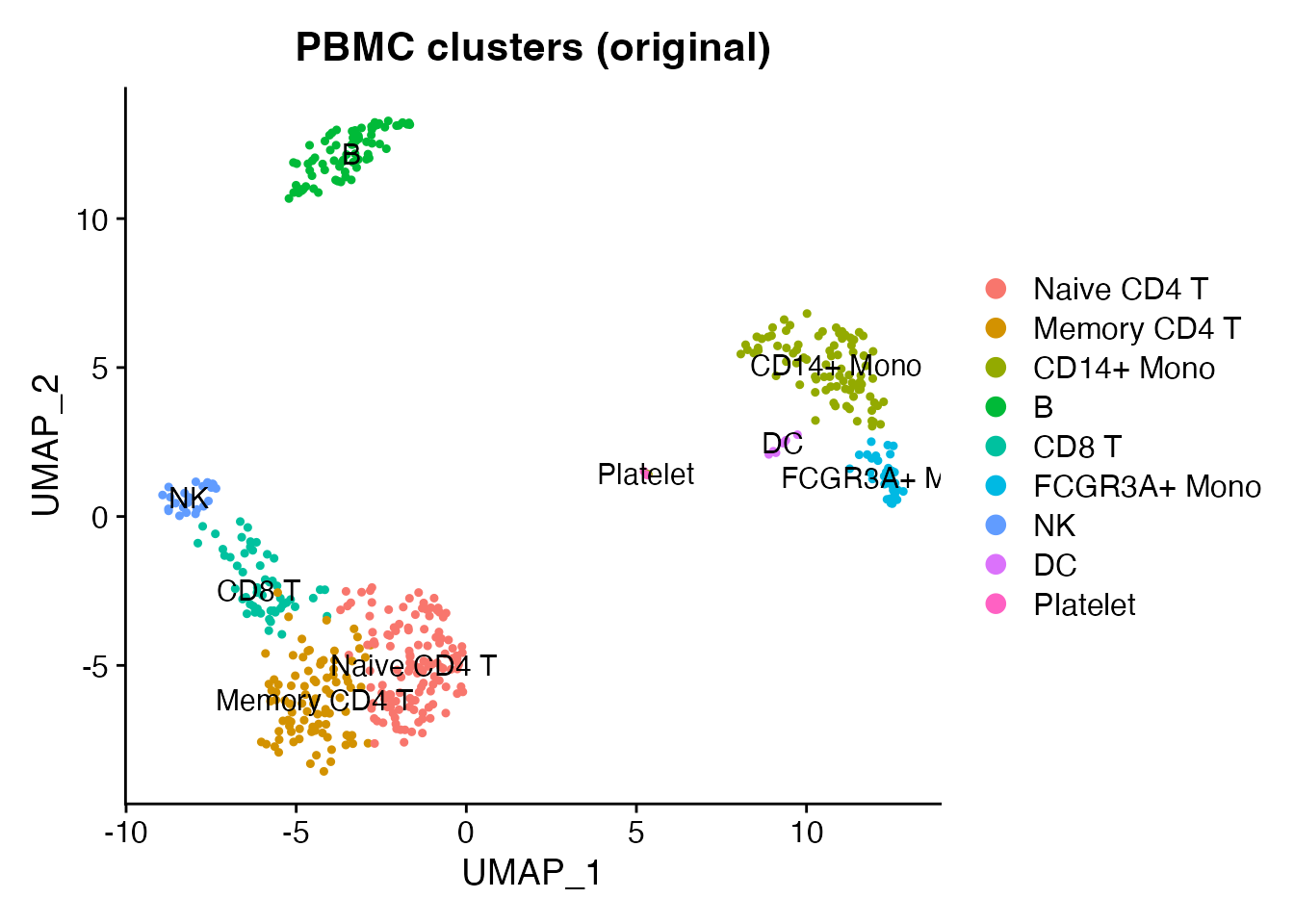
Roundtrip through h5ad
Write the Seurat object to h5ad and read it back.
h5ad_path <- file.path(tempdir(), "pbmc_atac_demo.h5ad")
writeH5AD(obj, h5ad_path, overwrite = TRUE)
cat("Wrote:", basename(h5ad_path), "\n")
#> Wrote: pbmc_atac_demo.h5ad
cat("File size:", round(file.size(h5ad_path) / 1024^2, 1), "MB\n")
#> File size: 0.9 MB
loaded <- readH5AD(h5ad_path)
cat("Loaded:", ncol(loaded), "cells x", nrow(loaded), "features\n")
#> Loaded: 500 cells x 2000 featuresVerify preservation
Dimensions and barcodes
cat("Cells match:", ncol(obj) == ncol(loaded), "\n")
#> Cells match: TRUE
cat("Features match:", nrow(obj) == nrow(loaded), "\n")
#> Features match: TRUE
orig_cells <- sort(colnames(obj))
rt_cells <- sort(colnames(loaded))
cat("Barcodes identical:", identical(orig_cells, rt_cells), "\n")
#> Barcodes identical: TRUE
orig_features <- sort(rownames(obj))
rt_features <- sort(rownames(loaded))
cat("Feature names identical:", identical(orig_features, rt_features), "\n")
#> Feature names identical: TRUECount matrix
common_cells <- intersect(colnames(obj), colnames(loaded))
common_feats <- intersect(rownames(obj), rownames(loaded))
orig_vals <- as.numeric(GetAssayData(obj, layer = "counts")[
head(common_feats, 200), head(common_cells, 200)])
rt_vals <- as.numeric(GetAssayData(loaded, layer = "counts")[
head(common_feats, 200), head(common_cells, 200)])
cat("Count values identical:", identical(orig_vals, rt_vals), "\n")
#> Count values identical: TRUEMetadata
shared_cols <- intersect(colnames(obj[[]]), colnames(loaded[[]]))
cat("Metadata columns preserved:", length(shared_cols), "/", ncol(obj[[]]), "\n")
#> Metadata columns preserved: 7 / 7
cat("Columns:", paste(shared_cols, collapse = ", "), "\n")
#> Columns: orig.ident, nCount_RNA, nFeature_RNA, seurat_annotations, percent.mt, RNA_snn_res.0.5, seurat_clusters
if ("seurat_annotations" %in% shared_cols) {
orig_ann <- as.character(obj$seurat_annotations[common_cells])
rt_ann <- as.character(loaded$seurat_annotations[common_cells])
cat("Cell annotations match:", identical(orig_ann, rt_ann), "\n")
}
#> Cell annotations match: TRUEEmbeddings
cat("Original reductions:", paste(names(obj@reductions), collapse = ", "), "\n")
#> Original reductions: pca, umap
cat("Loaded reductions:", paste(names(loaded@reductions), collapse = ", "), "\n")
#> Loaded reductions: pca, umap
if ("umap" %in% names(loaded@reductions)) {
orig_umap <- Embeddings(obj, "umap")[common_cells, ]
rt_umap <- Embeddings(loaded, "umap")[common_cells, ]
max_diff <- max(abs(orig_umap - rt_umap))
cat("UMAP max absolute difference:", max_diff, "\n")
}
#> UMAP max absolute difference: 0Compare plots
library(patchwork)
p1 <- DimPlot(obj, group.by = "seurat_annotations", label = TRUE, pt.size = 0.8) +
ggtitle("Original") + NoLegend()
p2 <- DimPlot(loaded, group.by = "seurat_annotations", label = TRUE, pt.size = 0.8) +
ggtitle("After h5ad roundtrip") + NoLegend()
p1 + p2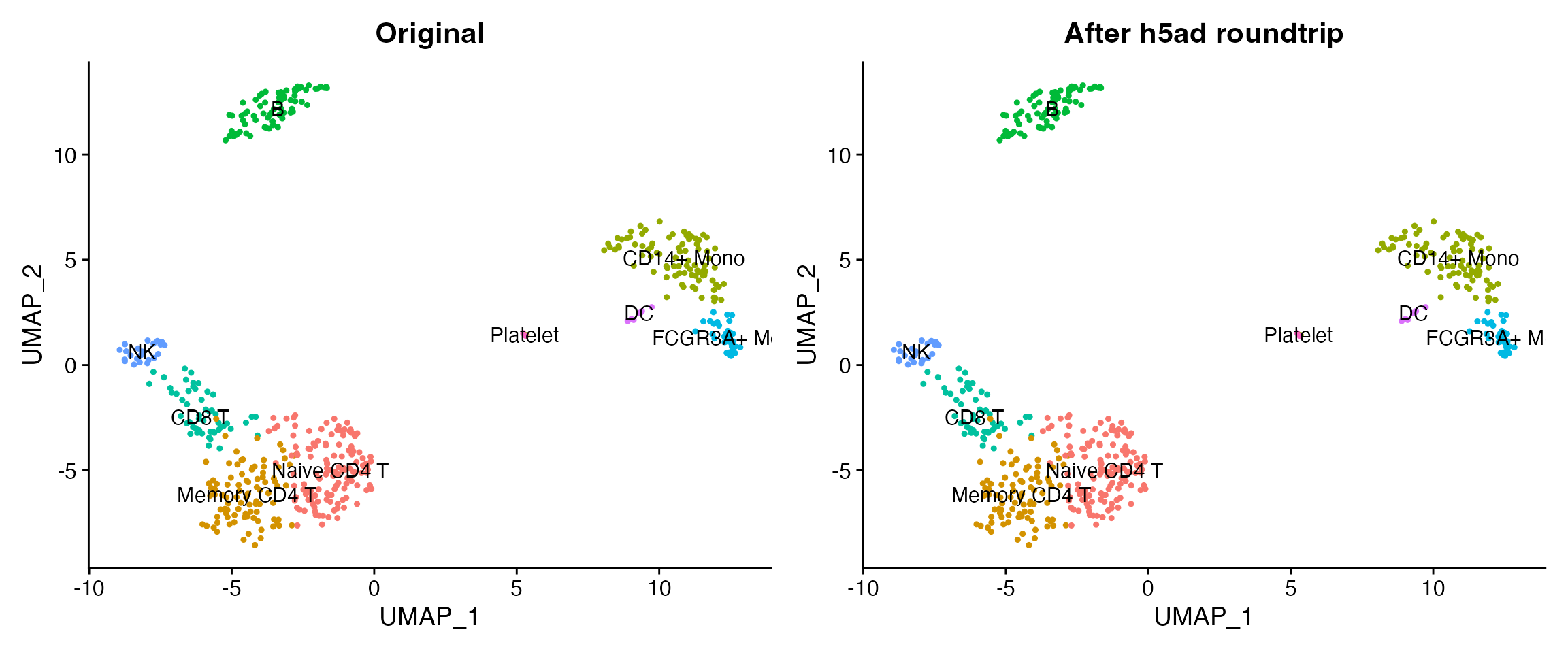
What is preserved vs. what needs separate handling
For real scATAC-seq data, scConvert preserves the core data but some Signac-specific components require separate handling:
Preserved by scConvert
| Component | Seurat/Signac | h5ad |
|---|---|---|
| Peak count matrix |
counts layer |
X |
| Cell metadata | meta.data |
obs |
| Peak metadata | meta.features |
var |
| LSI/PCA embeddings | reductions |
obsm |
| UMAP coordinates | reductions$umap |
obsm['X_umap'] |
| Cluster labels | meta.data$seurat_clusters |
obs['seurat_clusters'] |
Needs separate handling
| Component | Why | Workaround |
|---|---|---|
| Fragment files | Large tabix files, not part of h5ad | Copy fragments.tsv.gz + .tbi
separately |
| Gene annotations | GRanges object (R-specific) | Reload from EnsDb after import |
| Motif matrices | Signac-specific slot | Recompute with AddMotifs()
|
| ChromatinAssay class | R S4 class, not in h5ad | Upgrade with CreateChromatinAssay()
|
Upgrading to ChromatinAssay (if Signac is available)
After loading an h5ad file containing peak data, you can upgrade the
standard Seurat assay to a Signac ChromatinAssay for
peak-aware analyses.
library(Signac)
# Load h5ad containing peak-by-cell matrix
atac <- readH5AD("peaks.h5ad")
# Upgrade to ChromatinAssay
peak_counts <- GetAssayData(atac, layer = "counts")
atac[["peaks"]] <- CreateChromatinAssay(
counts = peak_counts,
sep = c("-", "-"),
min.cells = 0,
min.features = 0
)
# Re-attach annotations if needed
# library(EnsDb.Hsapiens.v86)
# Annotation(atac) <- GetGRangesFromEnsDb(ensdb = EnsDb.Hsapiens.v86)
# Fragments(atac) <- CreateFragmentObject("fragments.tsv.gz")Python interoperability
The exported h5ad is directly readable by scanpy or episcanpy.
# Requires Python: pip install scanpy
import scanpy as sc
adata = sc.read_h5ad("peaks.h5ad")
print(adata)
print(f"Peak names: {list(adata.var_names[:5])}")
# Standard scanpy pipeline works on peak matrices
sc.pp.normalize_total(adata)
sc.pp.log1p(adata)
sc.pp.pca(adata)
sc.pp.neighbors(adata)
sc.tl.umap(adata)
sc.pl.umap(adata, color="seurat_clusters")Clean up
unlink(h5ad_path)Session Info
sessionInfo()
#> R version 4.6.0 (2026-04-24)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] patchwork_1.3.2 ggplot2_4.0.3 Seurat_5.5.0 SeuratObject_5.4.0
#> [5] sp_2.2-1 scConvert_0.2.0
#>
#> loaded via a namespace (and not attached):
#> [1] RColorBrewer_1.1-3 jsonlite_2.0.0 magrittr_2.0.5
#> [4] spatstat.utils_3.2-2 farver_2.1.2 rmarkdown_2.31
#> [7] fs_2.1.0 ragg_1.5.2 vctrs_0.7.3
#> [10] ROCR_1.0-12 spatstat.explore_3.8-0 htmltools_0.5.9
#> [13] sass_0.4.10 sctransform_0.4.3 parallelly_1.47.0
#> [16] KernSmooth_2.23-26 bslib_0.10.0 htmlwidgets_1.6.4
#> [19] desc_1.4.3 ica_1.0-3 plyr_1.8.9
#> [22] plotly_4.12.0 zoo_1.8-15 cachem_1.1.0
#> [25] igraph_2.3.0 mime_0.13 lifecycle_1.0.5
#> [28] pkgconfig_2.0.3 Matrix_1.7-5 R6_2.6.1
#> [31] fastmap_1.2.0 MatrixGenerics_1.24.0 fitdistrplus_1.2-6
#> [34] future_1.70.0 shiny_1.13.0 digest_0.6.39
#> [37] S4Vectors_0.50.0 tensor_1.5.1 RSpectra_0.16-2
#> [40] irlba_2.3.7 GenomicRanges_1.64.0 textshaping_1.0.5
#> [43] labeling_0.4.3 progressr_0.19.0 spatstat.sparse_3.1-0
#> [46] httr_1.4.8 polyclip_1.10-7 abind_1.4-8
#> [49] compiler_4.6.0 bit64_4.8.0 withr_3.0.2
#> [52] S7_0.2.2 fastDummies_1.7.6 MASS_7.3-65
#> [55] tools_4.6.0 lmtest_0.9-40 otel_0.2.0
#> [58] httpuv_1.6.17 future.apply_1.20.2 goftest_1.2-3
#> [61] glue_1.8.1 nlme_3.1-169 promises_1.5.0
#> [64] grid_4.6.0 Rtsne_0.17 cluster_2.1.8.2
#> [67] reshape2_1.4.5 generics_0.1.4 hdf5r_1.3.12
#> [70] gtable_0.3.6 spatstat.data_3.1-9 tidyr_1.3.2
#> [73] data.table_1.18.2.1 BiocGenerics_0.58.0 BPCells_0.3.1
#> [76] spatstat.geom_3.7-3 RcppAnnoy_0.0.23 ggrepel_0.9.8
#> [79] RANN_2.6.2 pillar_1.11.1 stringr_1.6.0
#> [82] spam_2.11-3 RcppHNSW_0.6.0 later_1.4.8
#> [85] splines_4.6.0 dplyr_1.2.1 lattice_0.22-9
#> [88] survival_3.8-6 bit_4.6.0 deldir_2.0-4
#> [91] tidyselect_1.2.1 miniUI_0.1.2 pbapply_1.7-4
#> [94] knitr_1.51 gridExtra_2.3 Seqinfo_1.2.0
#> [97] IRanges_2.46.0 scattermore_1.2 stats4_4.6.0
#> [100] xfun_0.57 matrixStats_1.5.0 stringi_1.8.7
#> [103] lazyeval_0.2.3 yaml_2.3.12 evaluate_1.0.5
#> [106] codetools_0.2-20 tibble_3.3.1 cli_3.6.6
#> [109] uwot_0.2.4 xtable_1.8-8 reticulate_1.46.0
#> [112] systemfonts_1.3.2 jquerylib_0.1.4 Rcpp_1.1.1-1.1
#> [115] globals_0.19.1 spatstat.random_3.4-5 png_0.1-9
#> [118] spatstat.univar_3.1-7 parallel_4.6.0 pkgdown_2.2.0
#> [121] dotCall64_1.2 listenv_0.10.1 viridisLite_0.4.3
#> [124] scales_1.4.0 ggridges_0.5.7 purrr_1.2.2
#> [127] crayon_1.5.3 rlang_1.2.0 cowplot_1.2.0