Introduction
Zarr is a directory-based format for chunked, compressed arrays. It is widely used in cloud-native single-cell workflows – including CELLxGENE Census and SpatialData – because each chunk is an independent file that can be read in parallel from object stores like S3 or GCS. scConvert reads and writes Zarr v2 stores following the AnnData on-disk specification, with no Python dependency.
Read a Zarr store
We start with the shipped PBMC h5ad demo and convert it to Zarr so we can demonstrate the Zarr read/write workflow. This dataset has 230 cells, 220 genes, PCA/UMAP embeddings, neighbor graphs, and 9 annotated cell types.
h5ad_path <- system.file("testdata", "pbmc_small.h5ad", package = "scConvert")
zarr_path <- file.path(tempdir(), "pbmc_demo.zarr")
scConvert(h5ad_path, zarr_path, overwrite = TRUE, verbose = FALSE)
pbmc <- readZarr(zarr_path, verbose = FALSE)
pbmc
#> An object of class Seurat
#> 2000 features across 214 samples within 1 assay
#> Active assay: RNA (2000 features, 2000 variable features)
#> 2 layers present: counts, data
#> 2 dimensional reductions calculated: pca, umap
DimPlot(pbmc, reduction = "umap", group.by = "seurat_annotations",
label = TRUE, pt.size = 0.8) +
ggtitle("PBMC data read from Zarr") + NoLegend()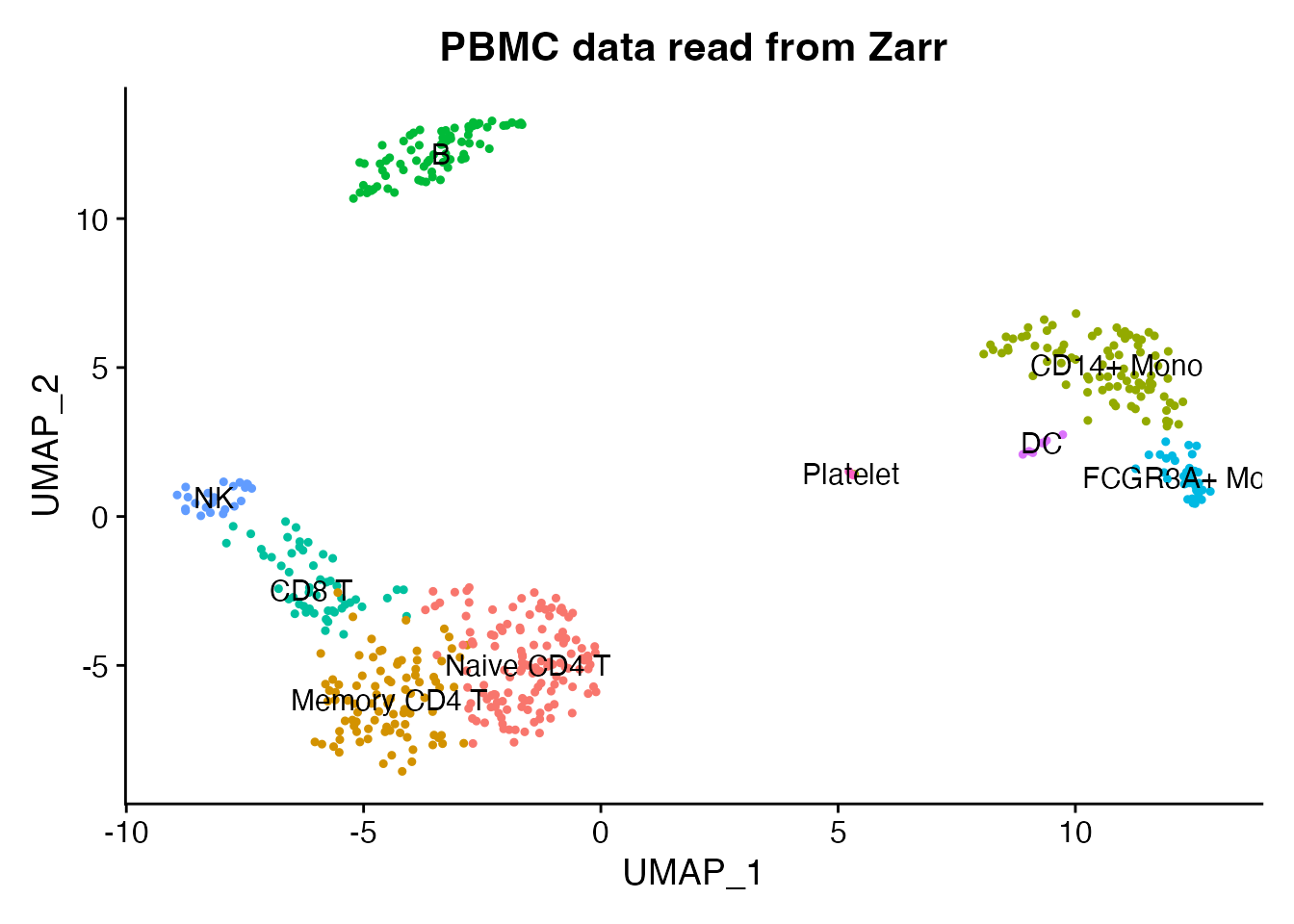
LYZ is a strong monocyte marker. We can verify that expression values were loaded correctly from the Zarr store.
FeaturePlot(pbmc, features = "LYZ", pt.size = 0.8) +
ggtitle("LYZ expression (from Zarr)")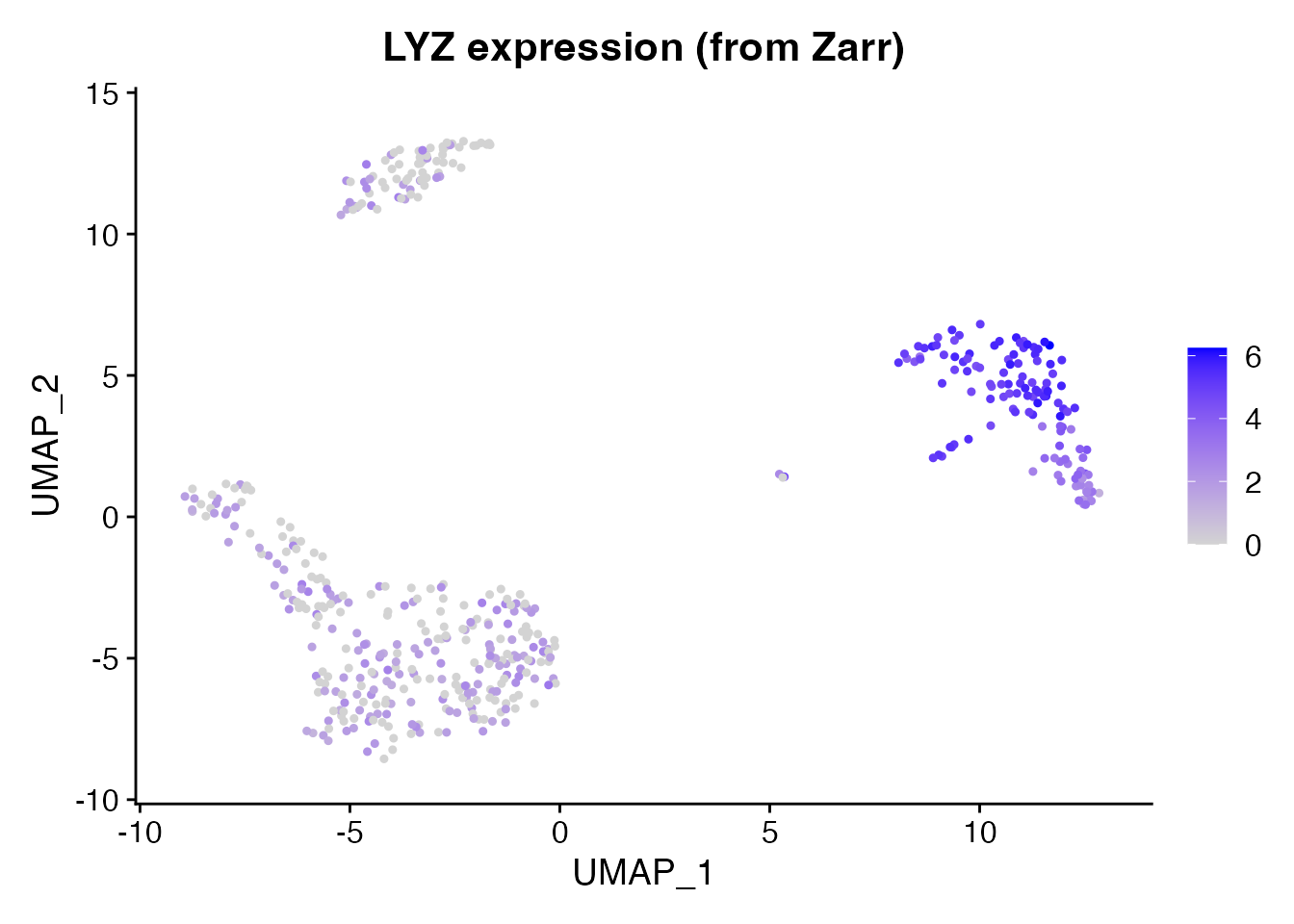
Write to Zarr and read back
Now we write the Seurat object to a new Zarr store and read it back to verify round-trip fidelity.
rt_path <- file.path(tempdir(), "pbmc_roundtrip.zarr")
writeZarr(pbmc, rt_path, overwrite = TRUE, verbose = FALSE)
pbmc_rt <- readZarr(rt_path, verbose = FALSE)
cat("Cells:", ncol(pbmc_rt), "| Genes:", nrow(pbmc_rt), "\n")
#> Cells: 214 | Genes: 2000
cat("Reductions:", paste(names(pbmc_rt@reductions), collapse = ", "), "\n")
#> Reductions: pca, umapVerify the round-trip
cat("Barcodes match:", identical(sort(Cells(pbmc)), sort(Cells(pbmc_rt))), "\n")
#> Barcodes match: TRUE
cat("Features match:", identical(sort(rownames(pbmc)), sort(rownames(pbmc_rt))), "\n")
#> Features match: TRUE
cat("Dimensions match:", identical(dim(pbmc), dim(pbmc_rt)), "\n")
#> Dimensions match: TRUEStreaming converters
For converting between file formats without loading data into R, scConvert provides streaming converters that copy fields directly between backends:
# h5ad <-> Zarr (no Seurat intermediate)
H5ADToZarr("data.h5ad", "data.zarr")
ZarrToH5AD("data.zarr", "data.h5ad")
# h5Seurat <-> Zarr
H5SeuratToZarr("data.h5seurat", "data.zarr")
ZarrToH5Seurat("data.zarr", "data.h5seurat")
# Or use the universal dispatcher
scConvert("data.h5ad", dest = "data.zarr", overwrite = TRUE)These are particularly useful for large datasets where materializing the full Seurat object would be expensive.
Python interoperability
Zarr stores produced by writeZarr() are directly
readable by Python’s anndata.read_zarr() and scanpy.
Requires Python with anndata installed.
Session Info
sessionInfo()
#> R version 4.6.0 (2026-04-24)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.3 Seurat_5.5.0 SeuratObject_5.4.0 sp_2.2-1
#> [5] scConvert_0.2.0
#>
#> loaded via a namespace (and not attached):
#> [1] deldir_2.0-4 pbapply_1.7-4 gridExtra_2.3
#> [4] rlang_1.2.0 magrittr_2.0.5 RcppAnnoy_0.0.23
#> [7] otel_0.2.0 spatstat.geom_3.7-3 matrixStats_1.5.0
#> [10] ggridges_0.5.7 compiler_4.6.0 png_0.1-9
#> [13] systemfonts_1.3.2 vctrs_0.7.3 reshape2_1.4.5
#> [16] hdf5r_1.3.12 stringr_1.6.0 crayon_1.5.3
#> [19] pkgconfig_2.0.3 fastmap_1.2.0 labeling_0.4.3
#> [22] promises_1.5.0 rmarkdown_2.31 ragg_1.5.2
#> [25] bit_4.6.0 purrr_1.2.2 xfun_0.57
#> [28] cachem_1.1.0 jsonlite_2.0.0 goftest_1.2-3
#> [31] later_1.4.8 spatstat.utils_3.2-2 irlba_2.3.7
#> [34] parallel_4.6.0 cluster_2.1.8.2 R6_2.6.1
#> [37] ica_1.0-3 spatstat.data_3.1-9 bslib_0.10.0
#> [40] stringi_1.8.7 RColorBrewer_1.1-3 reticulate_1.46.0
#> [43] spatstat.univar_3.1-7 parallelly_1.47.0 lmtest_0.9-40
#> [46] jquerylib_0.1.4 scattermore_1.2 Rcpp_1.1.1-1.1
#> [49] knitr_1.51 tensor_1.5.1 future.apply_1.20.2
#> [52] zoo_1.8-15 sctransform_0.4.3 httpuv_1.6.17
#> [55] Matrix_1.7-5 splines_4.6.0 igraph_2.3.0
#> [58] tidyselect_1.2.1 abind_1.4-8 yaml_2.3.12
#> [61] spatstat.random_3.4-5 spatstat.explore_3.8-0 codetools_0.2-20
#> [64] miniUI_0.1.2 listenv_0.10.1 plyr_1.8.9
#> [67] lattice_0.22-9 tibble_3.3.1 withr_3.0.2
#> [70] shiny_1.13.0 S7_0.2.2 ROCR_1.0-12
#> [73] evaluate_1.0.5 Rtsne_0.17 future_1.70.0
#> [76] fastDummies_1.7.6 desc_1.4.3 survival_3.8-6
#> [79] polyclip_1.10-7 fitdistrplus_1.2-6 pillar_1.11.1
#> [82] KernSmooth_2.23-26 plotly_4.12.0 generics_0.1.4
#> [85] RcppHNSW_0.6.0 scales_1.4.0 globals_0.19.1
#> [88] xtable_1.8-8 glue_1.8.1 lazyeval_0.2.3
#> [91] tools_4.6.0 data.table_1.18.2.1 RSpectra_0.16-2
#> [94] RANN_2.6.2 fs_2.1.0 dotCall64_1.2
#> [97] cowplot_1.2.0 grid_4.6.0 tidyr_1.3.2
#> [100] nlme_3.1-169 patchwork_1.3.2 cli_3.6.6
#> [103] spatstat.sparse_3.1-0 textshaping_1.0.5 spam_2.11-3
#> [106] viridisLite_0.4.3 dplyr_1.2.1 uwot_0.2.4
#> [109] gtable_0.3.6 sass_0.4.10 digest_0.6.39
#> [112] progressr_0.19.0 ggrepel_0.9.8 htmlwidgets_1.6.4
#> [115] farver_2.1.2 htmltools_0.5.9 pkgdown_2.2.0
#> [118] lifecycle_1.0.5 httr_1.4.8 mime_0.13
#> [121] bit64_4.8.0 MASS_7.3-65